 Resource Balance Analysis
Resource Balance Analysis
Main | What is RBA | Models | Software | Contact
RBA models
Below you can find RBA models for a number of organisms. RBA models are encoded in an XML format, which contains a large amount of information and is not designed to be human-readable. However, network structure, problem parameters, and simulation results can be exported and visualised as tables. To inspect the models and browse their contents, click on the pictures below. You will see an overview page from which you can download model and simulation results as table files. To display the various tables (describing compounds, reactions, proteins, and many other model components), you can click on the corresponding region of the overview graphics or use the dropdown menus on top of the page. The organism icon on top the page will bring you back to the model main page, the larger icon on the left brings you back to this page. See "HowTo" (in the dropdown menus) for details about browsing the models.
Click on pictures to browse model contents
|
Bacillus subtilis (wild type) (Goelzer et al. 2015) 
|
Escherichia coli (wild type) (Bulović et al. 2019) 
|
Escherichia coli (semi-autotrophic) (Bulović et al. 2019) 
|
|
Vibrio natriegens (Coppens et al. 2023) 
|
Cupriavidus necator (R. eutropha H16) (Jahn et al. 2021) 
|
Minimal cell model (central carbon metabolism) 
|
|
Minimal cell model (Molenaar et al. 2009) 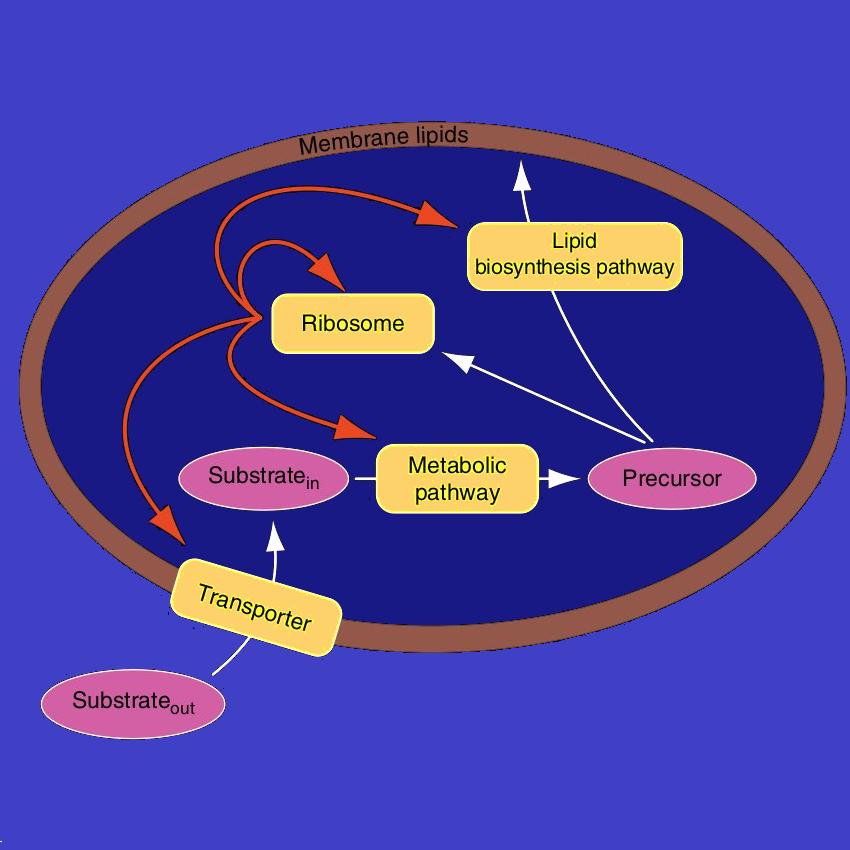
|
Download RBA models
These RBA models (in RBA XML format) can be downloaded from GitHub:
- Bacillus subtilis 168 (wild type)
- Escherichia coli K-12 (wild type)
- Escherichia coli K-12 (wild type), temperature dependent
- Escherichia coli K-12 (CO2-fixing engineered strain)
- Ralstonia eutropha H16 (also known as Cupriavidus necator)
- Vibrio natriegens (wild type)
- Arabidopsis thaliana Col-0 (generic plant cell)
- Arabidopsis thaliana Col-0 (generic plant cell), temperature dependent
Image copyright: Cupriavidus necator: courtesy of Professorship Microbial Biotechnology, TUM