 Resource Balance Analysis
Resource Balance Analysis
Main | What is RBA | Models | Software | Contact
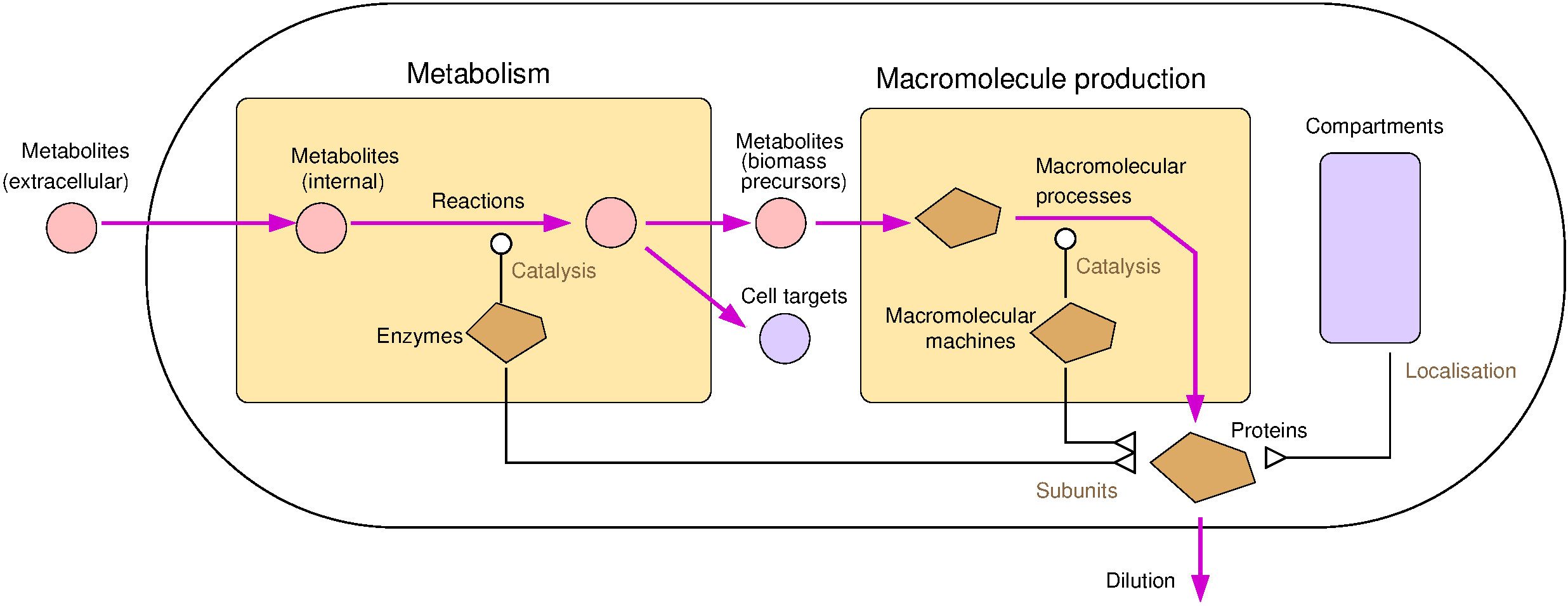
Resource balance analysis (RBA) is a framework for genome-scale models of cells and organisms. RBA models use a detailed blueprint of a cell to predict its precise macromolecular composition. Based on metabolic networks, models can be indefinitely refined by adding cellular processes (such as translation, transcription, secretion, or chaperoning). The RBA framework was originally developed to describe bacteria growing in batch cultures and has since then been extended to dynamical growth conditions, chemostat cultures, and eukaryotic organisms such as plants.
This website explains how to build and use RBA models and provides resources including models and python software. If you use our RBA code for your research, please cite our articles Bulović et al. (2019) (for RBApy) or Bodeit et al. (2023) (for RBAtools).